Loading and Defining Custom Spectral File Formats¶
Loading From a File¶
Specutils leverages the astropy io registry to provide an interface for conveniently
loading data from files. To create a custom loader, the user must define it in
a separate python file and place the file in their ~/.specutils directory.
Loading from a FITS File¶
A spectra with a Linear Wavelength Solution can be read using the read
method of the Spectrum1D class to parse the file name and
format
import os
from specutils import Spectrum1D
file_path = os.path.join('path/to/folder', 'file_with_1d_wcs.fits')
spec = Spectrum1D.read(file_path, format='wcs1d-fits')
This will create a Spectrum1D object that you can manipulate later.
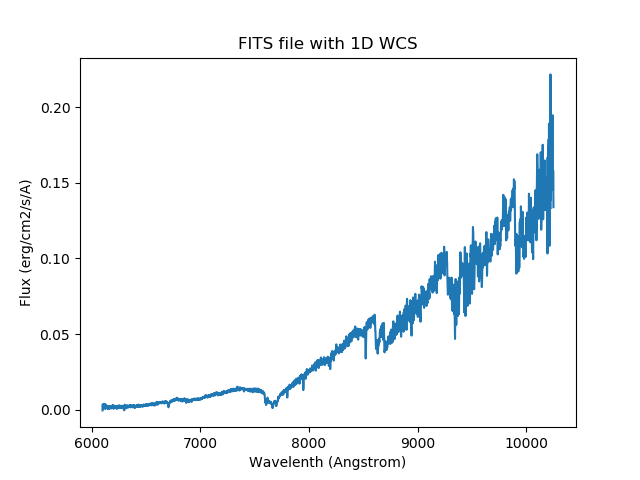
For instance, you could plot the spectrum.
import matplotlib.pyplot as plt
plt.title('FITS file with 1D WCS')
plt.xlabel('Wavelength (Angstrom)')
plt.ylabel('Flux (erg/cm2/s/A)')
plt.plot(spec.wavelength, spec.flux)
plt.show()
Creating a Custom Loader¶
Defining a custom loader consists of importing the
data_loader decorator from specutils and attaching
it to a function that knows how to parse the user’s data. The return object
of this function must be an instance of one of the spectral classes
(Spectrum1D, SpectrumCollection,
SpectrumList).
Optionally, the user may define an identifier function. This function acts to ensure that the data file being loaded is compatible with the loader function.
# ~/.specutils/my_custom_loader.py
import os
from astropy.io import fits
from astropy.nddata import StdDevUncertainty
from astropy.table import Table
from astropy.units import Unit
from astropy.wcs import WCS
from specutils.io.registers import data_loader
from specutils import Spectrum1D
# Define an optional identifier. If made specific enough, this circumvents the
# need to add ``format="my-format"`` in the ``Spectrum1D.read`` call.
def identify_generic_fits(origin, *args, **kwargs):
return (isinstance(args[0], str) and
os.path.splitext(args[0].lower())[1] == '.fits')
@data_loader("my-format", identifier=identify_generic_fits,
extensions=['fits'])
def generic_fits(file_name, **kwargs):
with fits.open(file_name, **kwargs) as hdulist:
header = hdulist[0].header
tab = Table.read(file_name)
meta = {'header': header}
wcs = WCS(hdulist[0].header)
uncertainty = StdDevUncertainty(tab["err"])
data = tab["flux"] * Unit("Jy")
return Spectrum1D(flux=data, wcs=wcs, uncertainty=uncertainty, meta=meta)
An extensions keyword can be provided. This allows for basic filename
extension matching in the case that the identifier function is not
provided.
It is possible to query the registry to return the list of loaders associated with a particular extension.
from specutils.io import get_loaders_by_extension
loaders = get_loaders_by_extension('fits')
The returned list contains the format labels that can be fed into the format
keyword argument of the Spectrum1D.read method.
After placing this python file in the user’s ~/.specutils directory, it
can be utilized by referencing its name in the read method of the
Spectrum1D class
from specutils import Spectrum1D
spec = Spectrum1D.read("path/to/data", format="my-format")
Loading Multiple Spectra¶
It is possible to create a loader that reads multiple spectra from the same
file. For the general case where none of the spectra are assumed to be the same
length, the loader should return a SpectrumList. Consider the
custom JWST data loader as an example:
import os
import glob
import warnings
from itertools import chain
import astropy.units as u
from astropy.units import Quantity
from astropy.table import Table
from astropy.io import fits
from astropy.nddata import StdDevUncertainty, VarianceUncertainty, InverseVariance
from astropy.time import Time
from astropy.wcs import WCS
from gwcs.wcstools import grid_from_bounding_box
from ...spectra import Spectrum1D, SpectrumList
from ..registers import data_loader
from ..parsing_utils import read_fileobj_or_hdulist
__all__ = ["jwst_x1d_single_loader", "jwst_x1d_multi_loader", "jwst_x1d_miri_mrs_loader"]
def identify_jwst_miri_mrs(origin, *args, **kwargs):
"""
Check whether the given set of files is a JWST MIRI MRS spectral data product.
"""
input = args[2]
# if string, it can be either a directory or a glob pattern (this last
# one not implemented yet due to astropy choking when passed an invalid
# file path string).
if isinstance(input, str):
if os.path.isdir(input):
return True
if len(glob.glob(input)) > 0:
return True
# or it can be either a list of file names, or a list of file objects
elif isinstance(input, (list, tuple)) and len(input) > 0:
return True
return False
def _identify_spec1d_fits(origin, extname, *args, **kwargs):
""" Generic spec 1d identifier function """
is_jwst = _identify_jwst_fits(*args)
with read_fileobj_or_hdulist(*args, memmap=False, **kwargs) as hdulist:
return (is_jwst and extname in hdulist and (extname, 2) not in hdulist)
def _identify_spec1d_multi_fits(origin, extname, *args, **kwargs):
"""
Check whether the given file is a JWST c1d/x1d spectral data product with many slits.
"""
is_jwst = _identify_jwst_fits(*args)
with read_fileobj_or_hdulist(*args, memmap=False, **kwargs) as hdulist:
return is_jwst and (extname, 2) in hdulist
def identify_jwst_c1d_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST c1d spectral data product.
"""
return _identify_spec1d_fits(origin, 'COMBINE1D', *args, **kwargs)
def identify_jwst_c1d_multi_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST c1d spectral data product with many slits.
"""
return _identify_spec1d_multi_fits(origin, 'COMBINE1D', *args, **kwargs)
def identify_jwst_x1d_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST x1d spectral data product.
"""
return _identify_spec1d_fits(origin, 'EXTRACT1D', *args, **kwargs)
def identify_jwst_x1d_multi_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST x1d spectral data product with many slits.
"""
return _identify_spec1d_multi_fits(origin, 'EXTRACT1D', *args, **kwargs)
def identify_jwst_s2d_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST s2d spectral data product.
"""
is_jwst = _identify_jwst_fits(*args)
with read_fileobj_or_hdulist(*args, memmap=False, **kwargs) as hdulist:
return (is_jwst and "SCI" in hdulist and ("SCI", 2) not in hdulist
and "EXTRACT1D" not in hdulist and len(hdulist["SCI"].data.shape) == 2)
def identify_jwst_s2d_multi_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST s2d spectral data product with many slits.
"""
is_jwst = _identify_jwst_fits(*args)
with read_fileobj_or_hdulist(*args, memmap=False, **kwargs) as hdulist:
return (is_jwst and ("SCI", 2) in hdulist and "EXTRACT1D" not in hdulist
and len(hdulist["SCI"].data.shape) == 2)
def identify_jwst_s3d_fits(origin, *args, **kwargs):
"""
Check whether the given file is a JWST s3d spectral data product.
"""
is_jwst = _identify_jwst_fits(*args)
with read_fileobj_or_hdulist(*args, memmap=False, **kwargs) as hdulist:
return (is_jwst and "SCI" in hdulist and "EXTRACT1D" not in hdulist
and len(hdulist["SCI"].data.shape) == 3)
def _identify_jwst_fits(*args):
"""
Check whether the given file is a JWST data product.
"""
try:
with read_fileobj_or_hdulist(*args, memmap=False) as hdulist:
return "ASDF" in hdulist and hdulist[0].header.get("TELESCOP") == "JWST"
# This probably means we didn't have a FITS file
except Exception:
return False
@data_loader(
"JWST c1d", identifier=identify_jwst_c1d_fits, dtype=Spectrum1D,
extensions=['fits'], priority=10,
)
def jwst_c1d_single_loader(file_obj, **kwargs):
"""
Loader for JWST c1d 1-D spectral data in FITS format
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
Spectrum1D
The spectrum contained in the file.
"""
spectrum_list = _jwst_spec1d_loader(file_obj, extname='COMBINE1D', **kwargs)
if len(spectrum_list) == 1:
return spectrum_list[0]
else:
raise RuntimeError(f"Input data has {len(spectrum_list)} spectra. "
"Use SpectrumList.read() instead.")
@data_loader(
"JWST c1d multi", identifier=identify_jwst_c1d_multi_fits,
dtype=SpectrumList, extensions=['fits'], priority=10,
)
def jwst_c1d_multi_loader(file_obj, **kwargs):
"""
Loader for JWST x1d 1-D spectral data in FITS format
Parameters
----------
file_obj: str, file-like, or HDUList
FITS file name, object (provided from name by Astropy I/O Registry),
or HDUList (as resulting from astropy.io.fits.open()).
Returns
-------
SpectrumList
A list of the spectra that are contained in the file.
"""
return _jwst_spec1d_loader(file_obj, extname='COMBINE1D', **kwargs)
@data_loader(
"JWST x1d", identifier=identify_jwst_x1d_fits, dtype=Spectrum1D,
extensions=['fits'], priority=10
)
def jwst_x1d_single_loader(file_obj, **kwargs):
"""
Loader for JWST x1d 1-D spectral data in FITS format
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
Spectrum1D
The spectrum contained in the file.
"""
spectrum_list = _jwst_spec1d_loader(file_obj, extname='EXTRACT1D', **kwargs)
if len(spectrum_list) == 1:
return spectrum_list[0]
else:
raise RuntimeError(f"Input data has {len(spectrum_list)} spectra. "
"Use SpectrumList.read() instead.")
@data_loader(
"JWST x1d multi", identifier=identify_jwst_x1d_multi_fits,
dtype=SpectrumList, extensions=['fits'], priority=10,
)
def jwst_x1d_multi_loader(file_obj, **kwargs):
"""
Loader for JWST x1d 1-D spectral data in FITS format
Parameters
----------
file_obj: str, file-like, or HDUList
FITS file name, object (provided from name by Astropy I/O Registry),
or HDUList (as resulting from astropy.io.fits.open()).
Returns
-------
SpectrumList
A list of the spectra that are contained in the file.
"""
return _jwst_spec1d_loader(file_obj, extname='EXTRACT1D', **kwargs)
@data_loader(
"JWST x1d MIRI MRS", identifier=identify_jwst_miri_mrs, dtype=SpectrumList,
extensions=['*'], priority=10,
)
def jwst_x1d_miri_mrs_loader(input, missing="raise", **kwargs):
"""
Loader for JWST x1d MIRI MRS spectral data in FITS format.
A single data set consists of a bunch of _x1d files corresponding to
a variety of wavelength bands. This reader reads them one by one and packs
the result into a SpectrumList instance.
Parameters
----------
input : list of str or file-like
List of FITS file names, or objects (provided from name by
Astropy I/O Registry). Alternatively, a directory path on
which glob.glob runs with pattern an implicit pattern "_x1d.fits",
or a directory path with a glob pattern already set.
missing : {'warn', 'silent'}
Allows the user to continue loading if one file is missing by setting
the value to "warn" or "silent". In the first case a warning will be issued
to the user, in the latter the file will silently be skipped. Any other
value will result in a FileNotFoundError if any files in the list are missing.
Returns
-------
SpectrumList
A list of the spectra that are contained in all the files.
"""
# If input is a list, go read each file. If directory, glob-expand
# list of file names.
if not isinstance(input, (list, tuple)):
if os.path.isdir(input):
file_list = glob.glob(os.path.join(input, "*_x1d.fits"), recursive=True)
else:
file_list = glob.glob(input, recursive=True)
else:
file_list = input
spectra = []
for file_obj in file_list:
try:
sp = _jwst_spec1d_loader(file_obj, **kwargs)
except FileNotFoundError as e:
if missing.lower() == "warn":
warnings.warn(f'Failed to load {file_obj}: {repr(e)}')
continue
elif missing.lower() == "silent":
continue
else:
raise FileNotFoundError(f"Failed to load {file_obj}: {repr(e)}. "
"To suppress this error, set argument missing='warn'")
spectra.append(sp)
# the call to `chain.from_iterable` allows us to handle multiple HDUs
# stored within multiple FITS files, all unpacked into one `SpectrumList`
return SpectrumList(chain.from_iterable(spectra))
def _jwst_spec1d_loader(file_obj, extname='EXTRACT1D', **kwargs):
"""Implementation of loader for JWST x1d 1-D spectral data in FITS format
Parameters
----------
file_obj: str, file-like, or HDUList
FITS file name, object (provided from name by Astropy I/O Registry),
or HDUList (as resulting from astropy.io.fits.open()).
extname: str
The name of the science extension. Either "COMBINE1D" or "EXTRACT1D". By default "EXTRACT1D".
Returns
-------
SpectrumList
A list of the spectra that are contained in the file.
"""
if extname not in ['COMBINE1D', 'EXTRACT1D']:
raise ValueError('Incorrect extname given for 1d spectral data.')
spectra = []
with read_fileobj_or_hdulist(file_obj, memmap=False, **kwargs) as hdulist:
primary_header = hdulist["PRIMARY"].header
for hdu in hdulist:
# Read only the BinaryTableHDUs named COMBINE1D/EXTRACT1D and SCI
if hdu.name != extname:
continue
# Correct some known bad unit strings before reading the table
bad_units = {"(MJy/sr)^2": "MJy2 sr-2"}
for c in hdu.columns:
if c.unit in bad_units:
c.unit = bad_units[c.unit]
data = Table.read(hdu)
wavelength = Quantity(data["WAVELENGTH"])
# Determine if FLUX or SURF_BRIGHT column should be returned
# based on whether it is point or extended source.
#
# SRCTYPE used to be in primary header, but was moved around. As
# per most recent pipeline definition, it should be in the
# EXTRACT1D extension.
#
# SRCTYPE should either be POINT or EXTENDED. In some cases, it is UNKNOWN
# or missing. If that's the case, default to using POINT as the SRCTYPE.
# Error out only when SRCTYPE is a bad value.
srctype = None
if "srctype" in hdu.header:
srctype = hdu.header.get("srctype", None)
# checking if SRCTPYE is missing or UNKNOWN
if not srctype or srctype == 'UNKNOWN':
warnings.warn('SRCTYPE is missing or UNKNOWN in JWST x1d loader. '
'Defaulting to srctype="POINT".')
srctype = 'POINT'
if srctype == "POINT":
flux = Quantity(data["FLUX"])
if 'ERROR' in data.colnames:
uncertainty = StdDevUncertainty(data["ERROR"])
elif 'FLUX_ERROR' in data.colnames:
uncertainty = StdDevUncertainty(data["FLUX_ERROR"])
else:
uncertainty = None
elif srctype == "EXTENDED":
flux = Quantity(data["SURF_BRIGHT"])
uncertainty = StdDevUncertainty(data["SB_ERROR"])
else:
raise RuntimeError(f"Keyword SRCTYPE is {srctype}. It should "
"be 'POINT' or 'EXTENDED'. Can't decide between `flux` and "
"`surf_bright` columns.")
# Merge primary and slit headers and dump into meta
slit_header = hdu.header
header = primary_header.copy()
header.extend(slit_header, strip=True, update=True)
meta = {'header': header}
spec = Spectrum1D(flux=flux, spectral_axis=wavelength,
uncertainty=uncertainty, meta=meta)
spectra.append(spec)
return SpectrumList(spectra)
@data_loader(
"JWST s2d", identifier=identify_jwst_s2d_fits, dtype=Spectrum1D,
extensions=['fits'], priority=10,
)
def jwst_s2d_single_loader(filename, **kwargs):
"""
Loader for JWST s2d 2D rectified spectral data in FITS format.
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
Spectrum1D
The spectrum contained in the file.
"""
spectrum_list = _jwst_s2d_loader(filename, **kwargs)
if len(spectrum_list) == 1:
return spectrum_list[0]
elif len(spectrum_list) > 1:
raise RuntimeError(f"Input data has {len(spectrum_list)} spectra. "
"Use SpectrumList.read() instead.")
else:
raise RuntimeError(f"Input data has {len(spectrum_list)} spectra.")
@data_loader(
"JWST s2d multi", identifier=identify_jwst_s2d_multi_fits,
dtype=SpectrumList, extensions=['fits'], priority=10,
)
def jwst_s2d_multi_loader(filename, **kwargs):
"""
Loader for JWST s2d 2D rectified spectral data in FITS format.
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
SpectrumList
The spectra contained in the file.
"""
return _jwst_s2d_loader(filename, **kwargs)
def _jwst_s2d_loader(filename, **kwargs):
"""
Loader for JWST s2d 2D rectified spectral data in FITS format.
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
SpectrumList
The spectra contained in the file.
"""
from stdatamodels import asdf_in_fits
spectra = []
slits = None
# Get a list of GWCS objects from the slits
with fits.open(filename, memmap=False) as hdulist, asdf_in_fits.open(hdulist) as af:
# Slits can be listed under "slits", "products" or "exposures"
if "products" in af.tree:
slits = "products"
elif "exposures" in af.tree:
slits = "exposures"
elif "slits" in af.tree:
slits = "slits"
else:
# No slits. Case for s3d cubes
wcslist = [af.tree["meta"]["wcs"]]
# Create list of the GWCS objects, one for each slit
if slits is not None:
wcslist = [slit["meta"]["wcs"] for slit in af.tree[slits]]
primary_header = hdulist["PRIMARY"].header
hdulist_sci = [hdu for hdu in hdulist if hdu.name == "SCI"]
for hdu, wcs in zip(hdulist_sci, wcslist):
# Get flux
try:
flux_unit = u.Unit(hdu.header["BUNIT"])
except (ValueError, KeyError):
flux_unit = None
# The dispersion axis is 1 or 2. 1=x, 2=y.
dispaxis = hdu.header.get("DISPAXIS")
if dispaxis is None:
dispaxis = hdulist["PRIMARY"].header.get("DISPAXIS")
# Get the wavelength array from the GWCS object which returns a
# tuple of (RA, Dec, lambda)
grid = grid_from_bounding_box(wcs.bounding_box)
_, _, lam = wcs(*grid)
_, _, lam_unit = wcs.output_frame.unit
# Make sure the dispersion axis is the same for every spatial axis
# for s2d data
if dispaxis == 1:
flux_array = hdu.data
wavelength_array = lam[0]
# Make sure all rows are the same
if not (lam == wavelength_array).all():
raise RuntimeError("This 2D or 3D spectrum is not rectified "
"and cannot be loaded into a Spectrum1D object.")
elif dispaxis == 2:
flux_array = hdu.data.T
wavelength_array = lam[:, 0]
# Make sure all columns are the same
if not (lam.T == lam[None, :, 0]).all():
raise RuntimeError("This 2D or 3D spectrum is not rectified "
"and cannot be loaded into a Spectrum1D object.")
else:
raise RuntimeError("This 2D spectrum has an unknown dispaxis "
"and cannot be loaded into a Spectrum1D object.")
flux = Quantity(flux_array, unit=flux_unit)
wavelength = Quantity(wavelength_array, unit=lam_unit)
# Merge primary and slit headers and dump into meta
slit_header = hdu.header
header = primary_header.copy()
header.extend(slit_header, strip=True, update=True)
meta = {'header': header}
spec = Spectrum1D(flux=flux, spectral_axis=wavelength, meta=meta)
spectra.append(spec)
return SpectrumList(spectra)
@data_loader(
"JWST s3d", identifier=identify_jwst_s3d_fits, dtype=Spectrum1D,
extensions=['fits'], priority=10,
)
def jwst_s3d_single_loader(filename, **kwargs):
"""
Loader for JWST s3d 3D rectified spectral data in FITS format.
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
Spectrum1D
The spectrum contained in the file.
"""
spectrum_list = _jwst_s3d_loader(filename, **kwargs)
if len(spectrum_list) == 1:
return spectrum_list[0]
elif len(spectrum_list) > 1:
raise RuntimeError(f"Input data has {len(spectrum_list)} spectra. "
"Use SpectrumList.read() instead.")
else:
raise RuntimeError(f"Input data has {len(spectrum_list)} spectra.")
def _jwst_s3d_loader(filename, **kwargs):
"""
Loader for JWST s3d 3D rectified spectral data in FITS format.
Parameters
----------
filename : str
The path to the FITS file
Returns
-------
SpectrumList
The spectra contained in the file.
"""
from stdatamodels import asdf_in_fits
spectra = []
# Get a list of GWCS objects from the slits
with fits.open(filename, memmap=False) as hdulist, asdf_in_fits.open(hdulist) as af:
wcslist = [af.tree["meta"]["wcs"]]
primary_header = hdulist["PRIMARY"].header
hdulist_sci = [hdu for hdu in hdulist if hdu.name == "SCI"]
for hdu, wcs in zip(hdulist_sci, wcslist):
# Get flux
try:
flux_unit = u.Unit(hdu.header["BUNIT"])
except (ValueError, KeyError):
flux_unit = None
# The spectral axis is first. We need it last
flux_array = hdu.data.T
flux = Quantity(flux_array, unit=flux_unit)
# Get the wavelength array from the GWCS object which returns a
# tuple of (RA, Dec, lambda).
# Since the spatial and spectral axes are orthogonal in s3d data,
# it is much faster to compute a slice down the spectral axis.
grid = grid_from_bounding_box(wcs.bounding_box)[:, :, 0, 0]
_, _, wavelength_array = wcs(*grid)
_, _, wavelength_unit = wcs.output_frame.unit
wavelength = Quantity(wavelength_array, unit=wavelength_unit)
# The GWCS is currently broken for some IFUs, here we work around that
wcs = None
if wavelength.shape[0] != flux.shape[-1]:
# Need MJD-OBS for this workaround
if 'MJD-OBS' not in hdu.header:
for key in ('MJD-BEG', 'DATE-OBS'): # Possible alternatives
if key in hdu.header:
if key.startswith('MJD'):
hdu.header['MJD-OBS'] = hdu.header[key]
break
else:
t = Time(hdu.header[key])
hdu.header['MJD-OBS'] = t.mjd
break
wcs = WCS(hdu.header)
# Swap to match the flux transpose
wcs = wcs.swapaxes(-1, 0)
# Merge primary and slit headers and dump into meta
slit_header = hdu.header
header = primary_header.copy()
header.extend(slit_header, strip=True, update=True)
meta = {'header': header}
# get uncertainty information
ext_name = primary_header.get("ERREXT", "ERR")
err_type = hdulist[ext_name].header.get("ERRTYPE", 'ERR')
err_unit = hdulist[ext_name].header.get("BUNIT", None)
err_array = hdulist[ext_name].data.T
# ERRTYPE can be one of "ERR", "IERR", "VAR", "IVAR"
# but mostly ERR for JWST cubes
# see https://jwst-pipeline.readthedocs.io/en/latest/jwst/data_products/science_products.html#s3d
if err_type == "ERR":
err = StdDevUncertainty(err_array, unit=err_unit)
elif err_type == 'VAR':
err = VarianceUncertainty(err_array, unit=err_unit)
elif err_type == 'IVAR':
err = InverseVariance(err_array, unit=err_unit)
elif err_type == 'IERR':
warnings.warn("Inverse error is not yet a supported astropy.nddata "
"uncertainty. Setting err to None.")
err = None
# get mask information
mask_name = primary_header.get("MASKEXT", "DQ")
mask = hdulist[mask_name].data.T
if wcs is not None:
spec = Spectrum1D(flux=flux, wcs=wcs, meta=meta, uncertainty=err, mask=mask)
else:
spec = Spectrum1D(flux=flux, spectral_axis=wavelength, meta=meta,
uncertainty=err, mask=mask)
spectra.append(spec)
return SpectrumList(spectra)
Note that by default, any loader that uses dtype=Spectrum1D will also
automatically add a reader for SpectrumList. This enables user
code to call specutils.SpectrumList.read in
all cases if it can’t make assumptions about whether a loader returns one or
many Spectrum1D objects. This method is available since
SpectrumList makes use of the Astropy IO registry (see
astropy.io.registry.read).
Creating a Custom Writer¶
Similar to creating a custom loader, a custom data writer may also be defined.
This again will be done in a separate python file and placed in the user’s
~/.specutils directory to be loaded into the astropy io registry.
# ~/.spectacle/my_writer.py
from astropy.table import Table
from specutils.io.registers import custom_writer
@custom_writer("fits-writer")
def generic_fits(spectrum, file_name, **kwargs):
flux = spectrum.flux.value
disp = spectrum.spectral_axis.value
meta = spectrum.meta
tab = Table([disp, flux], names=("spectral_axis", "flux"), meta=meta)
tab.write(file_name, format="fits")
The custom writer can be used by passing the name of the custom writer to the
format argument of the write method on the
Spectrum1D.
spec = Spectrum1D(flux=np.random.sample(100) * u.Jy,
spectral_axis=np.arange(100) * u.AA)
spec.write("my_output.fits", format="fits-writer")
Reference/API¶
A module containing the mechanics of the specutils io registry.
Functions¶
|
Wraps a function that can be added to an |
|
|
|
Retrieve a list of loader labels associated with a given extension. |
|
Attempt to identify a spectrum file format |